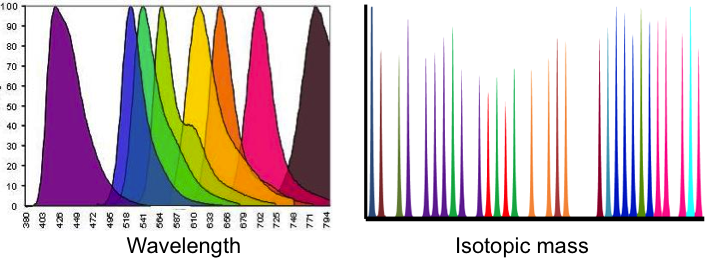
Mass cytometry is a single-cell analysis technology that overcomes most of the limitations of traditional flow cytometry. Currently, the number of reporters that can be measured by fluorescence-based flow cytometry is restricted by spectral overlap to a maximum of about ten. Since many of these ten colors would be used to stain surface markers for cell typing, very few colors remain available to measure other parameters such as phosphorylation or quantity of cellular proteins.
The CyTOF (Cytology Time of Flight) is a newly developed hybrid instrument that combines flow cytometry with mass spectrometry to allow single cell mass spectrometry using stable elemental mass reporters. This technique affords, at least, a 10-fold increase in the number of parameters that can be measured simultaneously. This advance is accompanied by the development of new software tools, such as Spanning-tree Progression Analysis of Density normalized Event (SPADE analysis), that allow the visualization of this high-dimension data. The CyTOF Flow Core will apply this technology to the phenotypic analysis of genetic perturbations generated in the Projects.
Aim 1. Create and standardize antibody and single-cell mRNA expression panels that cover all major surface markers and intracellular signaling in immune subsets relevant for the biology being interrogated in Projects 1-3
Develop perturbation conditions using cytokines and innate immunity perturbation conditions for robotic high-throughput analyses. Initiate creation of and testing of RNA probes for multiple cytokines and chemokines to be run alongside surface panels and intracellular signaling response panels.
Aim 2. Work directly with projects to enable full detailing of 2 mutant mouse strains per month, entailing baseline perturbation conditions as well as specific TLR responses and pathogen testing
Three CyTOF units are installed at Stanford University-two in the Nolan laboratory and one in the Human Immune Monitoring Core (under the direction of Mark Davis). Two more instruments are currently being set for acquisition by Stanford given what we believe to be the potential of this instrument to dramatically change the manner in which single cell studies can be done-especially that in the immune system. All of these instruments, and the techniques mentioned above, as well as a defining lead in computational ability in this new field (being developed at Stanford) are available for application in this Core. The Project sites will choose mutant strains for analysis. We will work with the Projects to determine appropriate signaling systems to be studied, cell types to focus upon, and stimulation conditions. Where pathogen testing at Stanford is to be undertaken (murine ectromelia for instance) or where shipment of mouse strains is prohibitive (fragile strains, low fecundity), we will provide pre-configured stimulation conditions and train Projects personnel how to stimulate and fix cells at their own facilities for shipment to our site.
Aim 3. Undertake analyses of high parameter datasets (SPADE and other developed algorithms) and provision of results through CytoBank
We will use the mouse strains studied in this U19 to create a “mapping” of mutation-induced changes to development and response profiles. We will work closely with Project 3 and the Systems Biology Core to enable full integration of our efforts to create a Systems Immunology data center accessible to the Project and other associated U19s.